The collaboration between the Venice Museum and UC Berkeley is highlighted in this Press Release.
The University of California, Berkeley and the Venice Museum of Natural History in Italy are collaborating to extensively catalog a museum collection using DNA information. The project started in April 2006, and collection of DNA sequence data was completed in 2011. The Venice Museum hosts the largest and best preserved fungal collection in Italy with more than 25,000 samples, representing approximately 6,000 species of fungi including many rare specimens. A diverse representation of the Venice collection was sampled and sent to U.C Berkeley, where a portion of the genome (the ribosomal DNA internal transcribed spacer, or ITS, region) was sequenced and analyzed.
![]()
Don’t have Google Earth? Download it for Free
Fungi are microbes that can only be identified when they produce fruiting bodies such as mushrooms; because mushrooms are only produced seasonally and sometimes very rarely, e.g. only once every several years, the resulting database will facilitate identification of fungi present in plants, in the soil, and in the air at all times, simply by comparing DNA information with the one generated by this and other studies. The DNA sequence data generated by this project are now available to the public and research communities and can enhance the ability to identify fungi in the environment. DNA-based identification of fungi in the environment can aid in the diagnosis of plant and animal diseases, enhance studies of fungal diversity and evolution, and lead to a better understanding of factors influencing nutrient cycling and productivity in forests and agricultural ecosystems.
The Venice collection is one of the first ones in the world to be extensively targeted for cataloging DNA information. The collection is unique as it is entirely generated and curated by volunteers, and in fact it is the outlet of the largest amateur mycological association in the world, the Associazione Micologica Bresadola, which currently has 13,000 active members. This ambitious project is made possible thanks to a collaborative effort between the Museums of the City of Venice and the Forest Pathology and Mycology Laboratory of the College of Natural Resources at U.C. Berkeley. The lead principal investigator is Dr. Matteo Garbelotto of U.C. Berkeley, aided by the Italian amateur mycologist Giovanni Robich and by the vice-director of the Venice Museum of Natural History, Dr. Luca Mizzan.
The scientific publication describing data collection and analyses in PLOSone:
Supplemental data from the project are available using the following links:
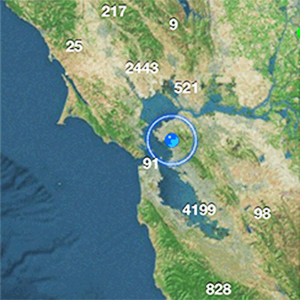
- Interactive map showing collection locations for each specimen from which a full-length ITS sequence was obtained (Google Earth .kmz file; must have Google Earth installed on your computer to view).
- List of specimens with full-length, double-stranded ITS sequences analyzed in the PLoS ONE publication (Supplemental Table S1; .xlsx format)
- Sequence data analyzed in the PLoS ONE publication, grouped by genus (zip archive). Please read the important readme file first!
- Contact Matteo Garbelotto – email matteo at matteolab.org to request access to additional, mostly single-stranded, sequence data generated during the project (see list)
PSATHYRELLA PROJECT AT THE VENICE MUSEUM OF NATURAL HISTORY
Download a partial list including published papers based on sequences generated by the collaboration between UC Berkeley and the Venice Museum of Natural History Link
Citizen Scientists: As Good As Gold
Papers Published Using Garbelotto’s Venice Sequences
Additional Links
Project Announcement
Press Release 12/13/06
Museum Herbarium Catalog