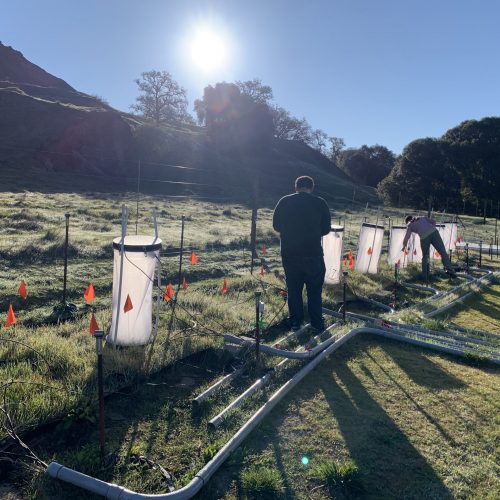
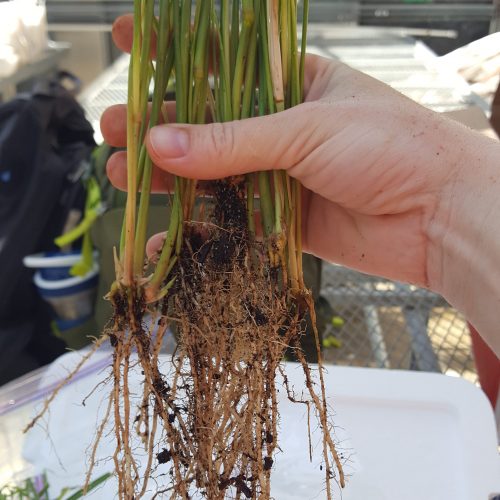
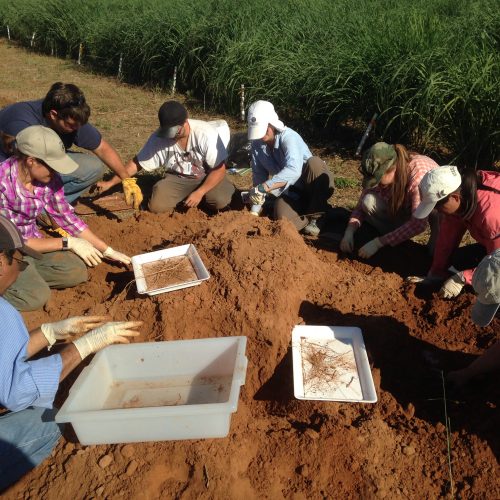
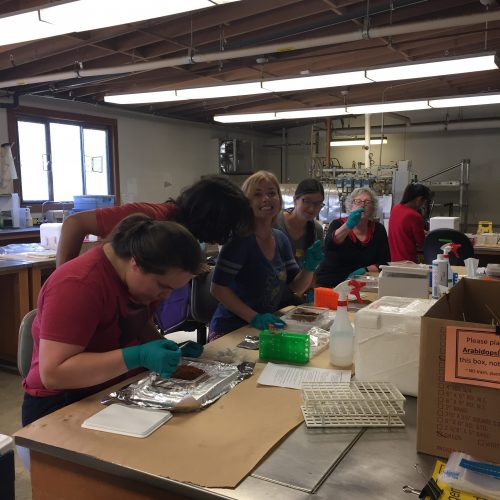
Next time you are outside, we encourage you to look down at the soil beneath your feet. Now, imagine scooping up a small thimble-full of that soil. Did you know that in just that tiny thimble of soil there are 1 billion bacteria and 1 million fungi? Amazing, right? We think so. Our lab studies soil microbial ecology – examining how all those microbes interact with growing plants and with each other, how they break down carbon substrates and cycle nutrients, and how they impact exchange of gases such as CO2, CH4, and N2O between the soil and the atmosphere.
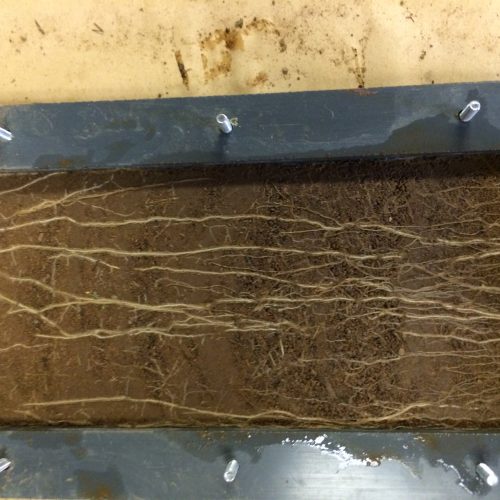
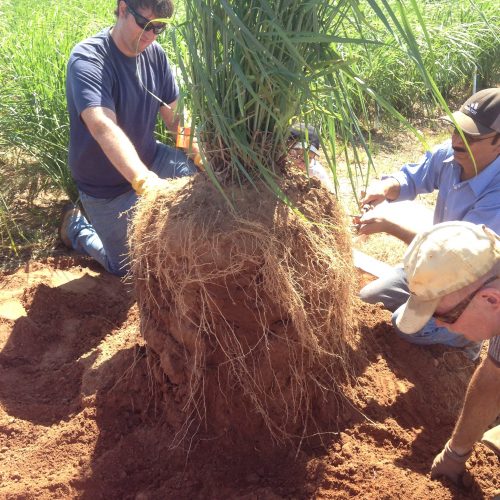
Our recent projects examine topics ranging ranging from: Microbial mediation of soil carbon dynamics; processes in the zone of soil around the plant root, called the rhizospere; using meta-omic approaches to link identity with function (metagenomes to metatranscriptomes to metaproteomes and metabolomes; digging deeper to examine microbial communities in saprolite (proto-soil); and microbial mediation of nitrogen cycling, with a particular focus on the highly reactive nitrous oxide.
Welcome to the Firestone Lab.
Latest Blog Posts
Recent PhD graduate Rachel Neurath starts postdoc at Lawrence Berkeley National Lab
Dr. Rachel Neurath will continue her natural science investigations and aspirations in...
Read More "Recent PhD graduate Rachel Neurath starts postdoc at Lawrence Berkeley National Lab"
Farewell to Jack Hagen
Our lab technician Jack Hagen has left the Firestone Lab for a...
Read More "Farewell to Jack Hagen"
A Firestone Farewell for Sarah & Nameer Baker
Nameer, Sarah and Naila at their farewell picnic (May 2020) Sarah and...
Read More "A Firestone Farewell for Sarah & Nameer Baker"
2020 Firestone Graduates
Photo of virtual Sather Gate from Blockley University We are excited to...
Read More "2020 Firestone Graduates"